Anti-Crossover Helicases

Several DNA helicases - enzymes that unwind DNA helices - have been implicated in important anti-crossover functions:
Biallelic pathogenic variants in BLM cause in Bloom syndrome. Clinical features of this disorder include small size and elevated incidence of a broad range of cancers. A predominant cellular feature is increased crossovers between sister chromatids, homologous chromosomes, and heterolgous chromosomes.
FANCM is one many genes in which pathogenic variants can cause Fanconi anemia. FANCM is thought to be important in recognizing interstrand DNA crosslinks and recruiting both the FA complex and BLM.
Inherited variants in SMARCAL1 cause Schimke immunoosseous dysplasia (SIOD), a rare form of dwarfism. The SMARCAL1 protein has been called an "annealing helicase" because it seems to anneal DNA more than unwind it.

Some mitotic crossovers arise during DSB repair. The synthesis-dependent strand anneling (SDSA) model (see full animation) explains hows DSBs can be repaired to give non-crossover products. One key step in this pathway is the disruption of the D-loop after repair DNA synthesis (right). Our studies of mutants defective in Blm, which encodes Drosophila BLM (DmBLM) led us to propose that it is the disruptase that promotes SDSA. This activity has also been well documented in vitro, and has been suggested to account for the anti-recombination activity of E. coli RecQ and the ability of RecQ to prevent illegitimate recombination (Harmon 1998).
M.D. Adams, M. McVey, and J. Sekelsky (2003) Drosophila BLM in double-strand break repair by synthesis-dependent strand annealing. Science 299: 265-267.
M. McVey, J. LaRocque, M.D. Adams, and J. Sekelsky (2004) Formation of deletions during double-strand break repair in Drosophila DmBlm mutants occurs after strand invasion. Proc. Natl. Acad. Sci. USA 101: 15694-15699.
M. McVey, S. Andersen, Y Broze, and J. Sekelsky (2007) Multiple functions of the Drosophila Blm helicase in maintaining genome stability. Genetics 176: 1979-1992.
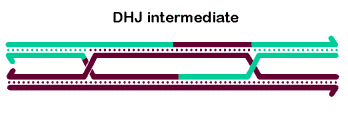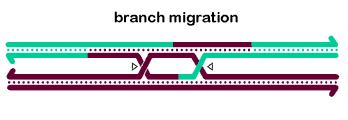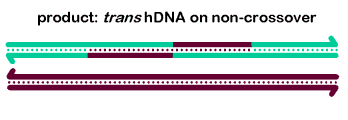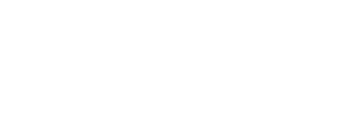
In vitro, BLM and associated proteins, including TOP3α, can carry out the dissolution of double-Holliday junction structures (Wu 2003, Plank 2006), as shown at right. The dHJ is thought to be a key intermediate in generating meiotic crossovers, at least in some organisms. Thus, many researches believe the dissolvase activity of BLM explains the ability of this protein to prevent mitotic crossovers. We believe it is doubtful that dHJ intermediates are common during DSB repair in cycling cells (SDSA is probably more common), but it is likely that dHJs do form during some other processes, such as some types of replication fork repair or when resolution is blocked during meiotic recombination.
S.L. Andersen and J. Sekelsky (2010)
Meiotic versus mitotic recombination: Two different routes for double-strand break repair.
Bioessays 32: 1058-1056.
BLM helicase has long been thought to play important roles in responding to blocked or broken replication forks. It is likely that the high frequency of mitotic crossing over in Drosophila Blm mutants (and in human cells lacking BLM) is a consequence of the inability of these mutants to properly manage blocked replication forks, leading to recombinogenic DSBs. We proposed that DSBs are generated at blocked forks by a class of structure-selective endonuclease we will refer to as resolvases. These resolvases typically cut fork structures only when helicases are unavailable to solve problems by catalyzing fork migration. Our models, which borrow heavily from models published by many other researchers, are based in part on the finding that loss of DmBLM and any of three resolvases simultaneous is lethal. We are testing various predictions of our models.
K. Trowbridge, K.S. McKim, S.J. Brill, and J. Sekelsky (2007) Synthetic lethality of Drosophila in the absence of the MUS81 endonuclease and the DmBlm helicase is associated with elevated apoptosis. Genetics 176: 1993-2001.
S.L. Andersen, D.T. Bergstralh, K.P. Kohl, J.R. LaRocque, C.B. Moore, and J. Sekelsky (2009) Drosophila MUS312 and the vertebrate ortholog BTBD12 interact with DNA structure-specific endonucleases in DNA repair and recombination. Molecular Cell 35: 128-135.
S.L. Andersen, H.K. Kuo, D. Savukoski, M.H. Brodsky, and J. Sekelsky (2011) Three structure-selective endonucleases are essential in the absence of BLM helicase in Drosophila. PLoS Genetics 7: e1002315.

